Table Of Content
The maximum stability for the last five 3' bases of a left or right primer. This enables our new graphic display that offers enhanced overview for your template and primers. The purpose of this subreddit is to help you learn (not complete your last-minute homework), and our rules are designed to reinforce this.
Quantification of rare somatic single nucleotide variants by droplet digital PCR using SuperSelective primers Scientific ... - Nature.com
Quantification of rare somatic single nucleotide variants by droplet digital PCR using SuperSelective primers Scientific ....
Posted: Fri, 03 Nov 2023 07:00:00 GMT [source]
NEBuilder HiFi DNA Assembly & Gibson Assembly
Tm (product)- It measures the melting temperature of the PCR product. Solely living organisms utilize RNA primers while invitro involves DNA primers. However, DNA primers are much preferred due to varied reasons such as stability, easy storage, fewer enzymes required to initiate synthesis. Optimise your research and save time with high quality gene synthesis and molecular biology services.
Tips and Considerations for Sensitive PCR Assays
However, the BLAST program reports a statistical significance, called “expectation value”(E – value) for each alignment which is an indicator for finding the match by chance. E – values ≤ 0.01 convey the homologous sequences (Altschul et al., 1990)( Karlin et al., 1990). E- value measures for assessing potential biological relationships (Raymaekers M et al, 2009). Despite the fact that Insilco tools provide valuable feedback, the specificity of the qPCR assay using the designed primers and probes has to be validated empirically with direct experimental evidence (Bustin et al., 2009). Bisulfite PCR is a multi-step process that enables users to determine the methylation status of CpG sites within a PCR amplicon. The first step in bisulfite PCR is bisulfite conversion of the DNA.
Golden Gate Cloning Method of Primer Designing
The default Table of thermodynamic parameters is "SantaLucia 1998" and the default Salt correction formula is "SantaLucia 1998" as recommended by primer3 program. Primer is a short stretch of sequence that serves as an initiation point for DNA synthesis. There can be a set of primers (forward and reverse) with a sequence complementary to the template DNA -a point of initiation synthesis. One critical point to keep in mind is that, whether to include the stop codon or not in the primer. For example, if you want to add a tag (fluorescent protein or a short peptide etc.) to the C terminal of the protein, you need to remove the stop codon from the reverse primer to let the tag be fused during translation.
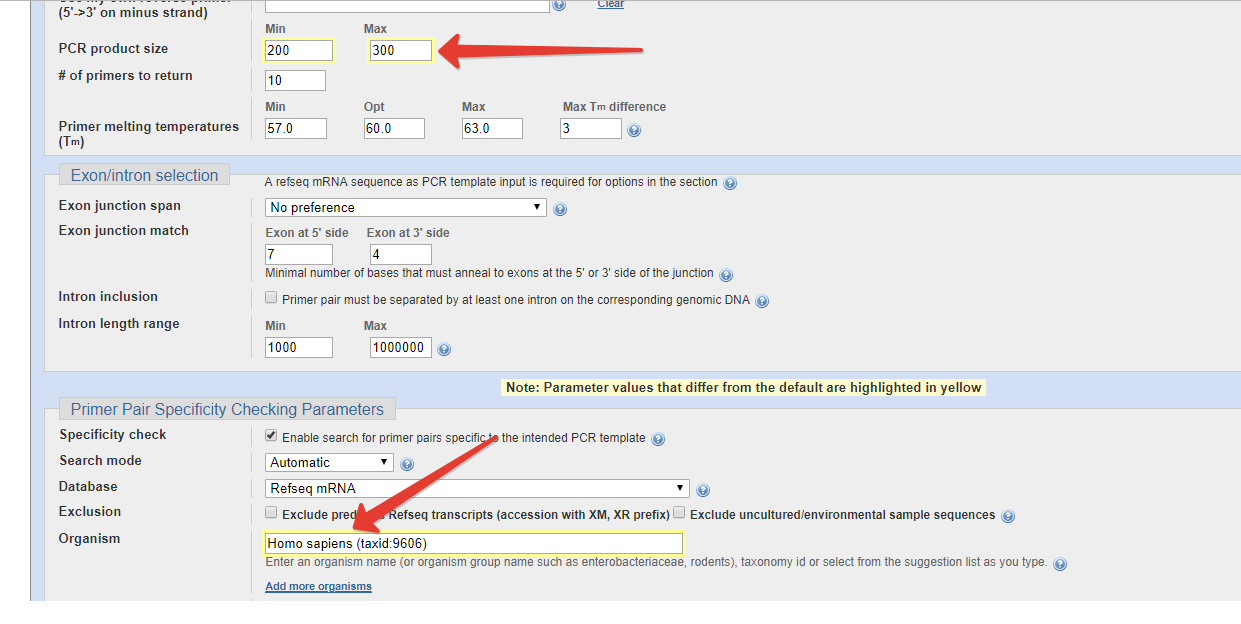
However, many downstream applications require extremely pure, concentrated DNA that is free of contaminants. In RT-qPCR, RNA is first converted into cDNA (reverse transcription, or RT), and then amplified with PCR as described in the previous section. During maturation, RNA is subjected to splicing, which removes introns from pre-mRNA sequences. For this reason, mRNA sequences differ from genomic DNA sequences. Thanks to the splicing it is possible to design a PCR assay specific to the RNA transcript. The maximum number of PCR targets (amplicons) to be shown when designing new primers.
Golden Gate Assembly
Cross-check if the sequence has been reverse complemented or not. Image showing the steps involved in reverse complementation in APE software. We are using APE (A Plasmid Editor) software to design the primers, which is free to use.
Choose a higher value if you need to perform more stringent search. This controls whether the primer should span an exon junction on your mRNA template. The option "Primer must span an exon-exon junction" will direct the program to return at least one primer (within a given primer pair) that spans an exon-exon junction. This is useful for limiting the amplification only to mRNA. You can also exclude such primers if you want to amplify mRNA as well as the corresponding genomic DNA.
The positions refer to the base numbers on the plus strand of your template (i.e., the "From" position should always be smaller than the "To" position for a given primer). Note that the position range of forward primer may not overlap with that of reverse primer. Although we have mentioned ideal conditions to make primers, it is not always possible to control the sequence selection (amplifying gene from the start codon for example).
Primer- Definition, Types, Primer Design Online Tools, Uses
If your goal is to sub-clone the gene or to express the protein without tags at the C-terminal of the gene, you will have to include the stop codon to prevent the translational fusion of vector sequences. In this demonstration, we are including the stop codon. To amplify any DNA sequence, two primers are necessary.

This specifies the range of total intron length on the corresponding genomic DNA that would separate the forward and revervse primers.
Polymerization direction is into the template stand rather outward of sequence as happens without reverse complementation. Methylation Specific PCR (MSP) relies on amplification to assess the methylation status at specific CpG sites after bisulfite conversion. Success with this system depends on the differential amplification of the template using methylated (M) and non-methylated (U) primer sets. While most of the considerations for MSP primer design are identical for those for bisulfite PCR, CpG sites within the primers are treated differently. Bisulfite PCR examines the methylation status of all the CpG sites present in a specific region, while MSP assesses the methylation at a single CpG site complementary to the 3’ end of the primer sequence. Enabling this option will make it much easier to find gene-specific primers since there is no need to distinguish between splice variants.
From the benefits of bonding primer to overcoming tight timelines and everything in between, get the guidance you need. Analyzing primer dimer formation is the primary important caution to be taken care of. However, it involves the determination of Gibbs free energy which aids to be the one. Although 5’ end was found to be more reliable than 3’ end. The comparison between DNA and RNA primers is listed below.